Link Found Between GRF genes in Sorghum and Aphid Resistance
Sorghum, the fifth most important cereal crop, provides food for more than a half billion people. It is also used as livestock feed and as a key component in biofuel. Biotic and abiotic stress resistant, sorghum is an ideal crop to be grown under adverse circumstances like poor soil or water stress conditions. Growth Regulating Factors (GRFs), which encode transcription factors regulating plant growth and development, also respond to changes in the environment. The study identified 8 Sorghum GRF genes containing QLQ (glutamine, leucine, glutamine) and WRC (tryptophan, arginine, cysteine) domains, which evolved in 4 clades. Understanding the role of the GRF gene family in pest resistance could contribute to the development of aphid resistant sorghum, which is of great interest to scientists, growers and breeders alike.
In an effort to better understand this role, Shi et al. from Shanghai Jiao Tong University, Hebei Academy of Agriculture & Forestry Sciences, Hebei Branch of China National Sorghum Improvement Center, Hebei Nijiao Brewing Technology Innovation Center, and Hebei Seed Management Station, identified eight GRF genes in the sorghum genome, evaluating their functional domain architecture and expression patterns under both normal and stressed (heat, salt, drought, aphid) conditions. These SbGRF genes were found to be expressed in most tissues and over half of them (including SbGRF1, 2, 3, 6, 7) expressed at the highest level in inflorescence. The expression level in inflorescence (flower clusters) is indicative of the role of GRFs in growth regulation. The authors hypothesize that flower development may be repressed by down-regulated SbGRFs and ectopic expression of the genes may increase the yield and biomass of sorghum plants.
The researchers performed a transcriptional analysis and discovered that more than half of the SbGRFs responded to abiotic stressors (heat, salinity and drought). This confirms the findings of other studies (Casadevall et al., 2013; Fina et al., 2017, Cao et al., 2020, Kim et al., 2012). However, the effects of biotic stressors, such as insects, on GRF expression has not been the focus of much study. The researchers found that the expression of SbGRF1, 2, 4 and 7 was greatly enhanced when sorghum plants were exposed to aphid stress. This indicates that GRFs help sorghum plants to defend against attacks from pests. The SbGRF response to aphids was validated by qRT-PCR. The results of this study could be used to target certain sorghum GRF genes for selective breeding in an effort to create aphid-resistant cultivars.
SorghumBase examples:
SorghumBase offers powerful comparative pangenomic views of extended gene families.
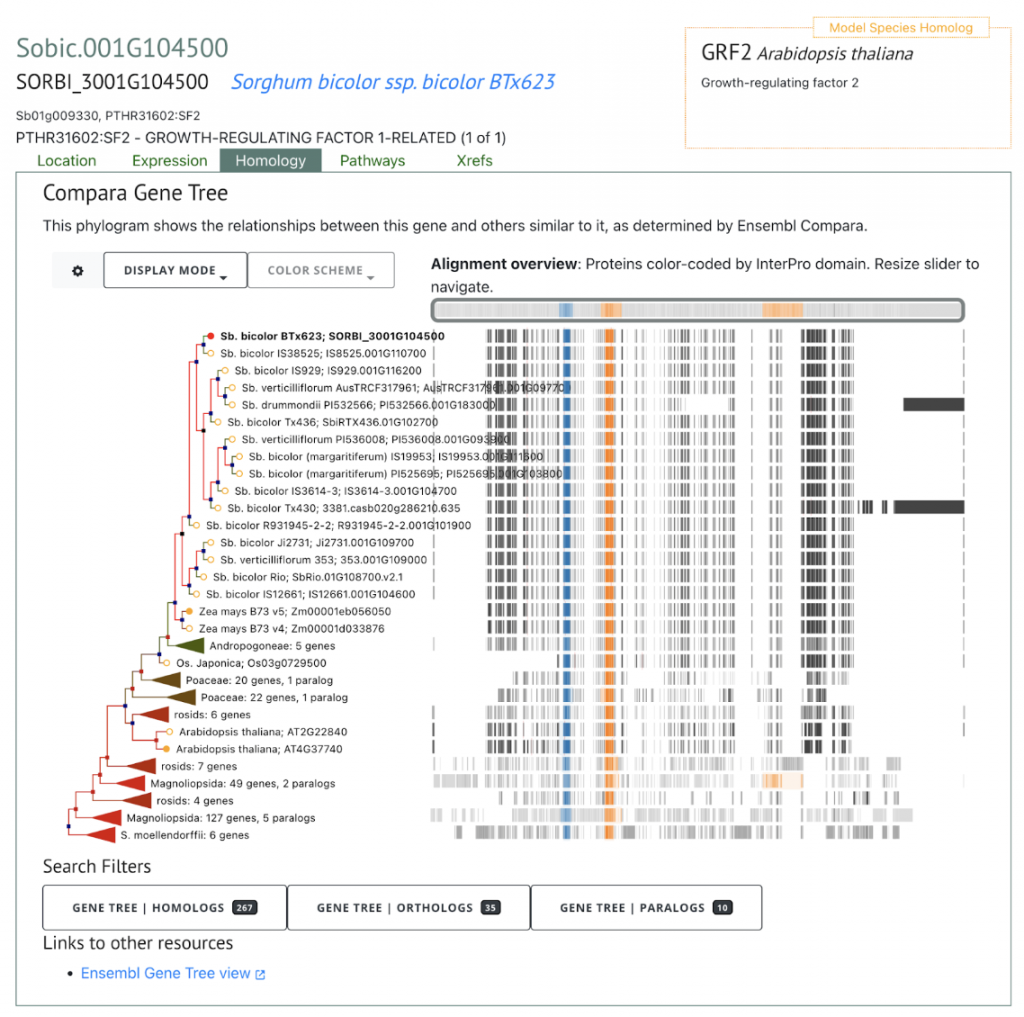
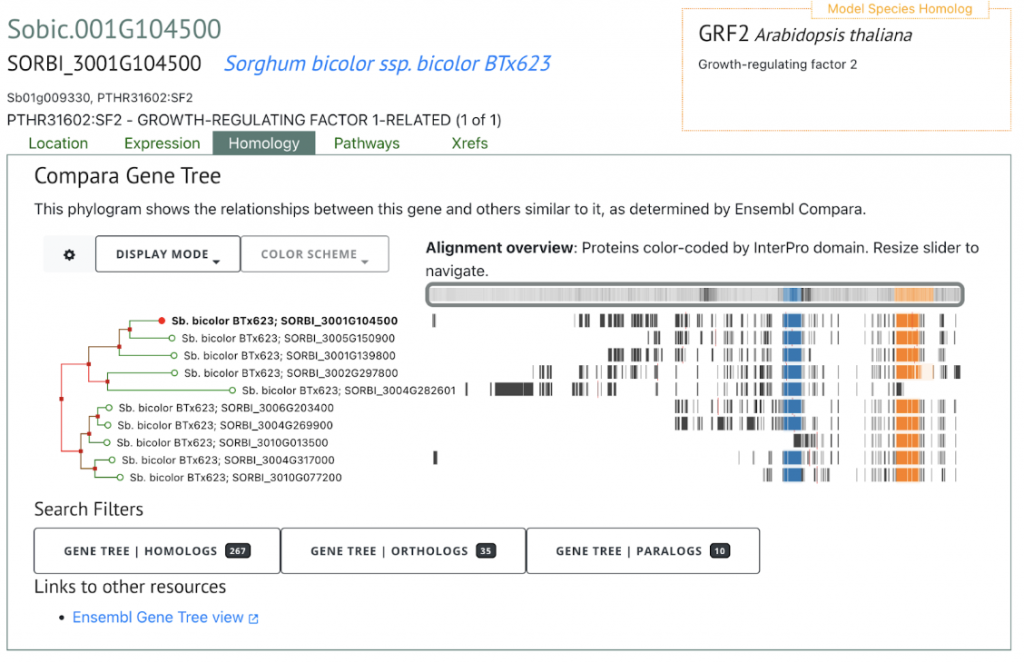

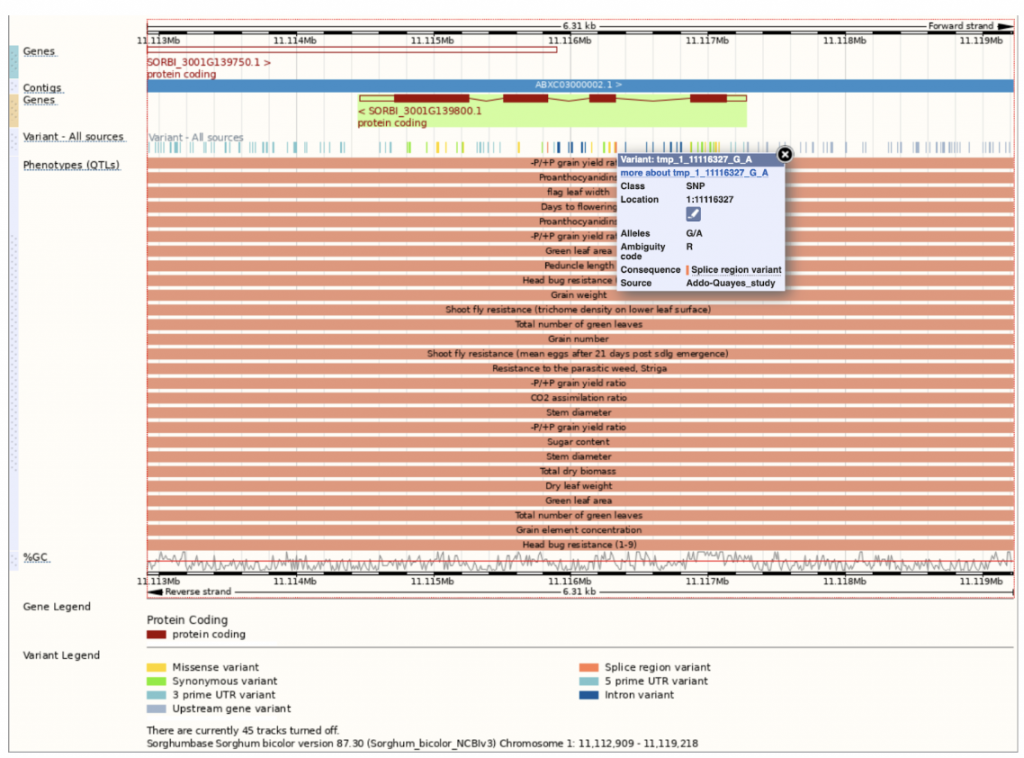
References
Cao JF, Huang JQ, Liu X, Huang CC, Zheng ZS, Zhang XF, Shangguan XX, Wang LJ, Zhang YG, Wendel JF, Grover CE, Chen ZW. Genome-wide characterization of the GRF family and their roles in response to salt stress in Gossypium. BMC Genomics. 2020 Aug 24;21(1):575. PMID: 32831017 DOI: 10.1186/s12864-020-06986-0. Read More
Casadevall R, Rodriguez RE, Debernardi JM, Palatnik JF, Casati P. Repression of growth regulating factors by the microRNA396 inhibits cell proliferation by UV-B radiation in Arabidopsis leaves. Plant Cell. 2013 Sep;25(9):3570-83. PMID: 24076976. DOI: 10.1105/tpc.113.117473. Read more
Fina J, Casadevall R, AbdElgawad H, Prinsen E, Markakis MN, Beemster GTS, Casati P. UV-B Inhibits Leaf Growth through Changes in Growth Regulating Factors and Gibberellin Levels. Plant Physiol. 2017 Jun;174(2):1110-1126.PMID: 28400494. DOI: 10.1104/pp.17.00365. Read more
Kim JS, Mizoi J, Kidokoro S, Maruyama K, Nakajima J, Nakashima K, Mitsuda N, Takiguchi Y, Ohme-Takagi M, Kondou Y, Yoshizumi T, Matsui M, Shinozaki K, Yamaguchi-Shinozaki K. Arabidopsis growth-regulating factor7 functions as a transcriptional repressor of abscisic acid- and osmotic stress-responsive genes, including DREB2A. Plant Cell. 2012 Aug;24(8):3393-405. PMID: 22942381. DOI: 10.1105/tpc.112.100933. Read more
Shi Y, Wang X, Wang J, Niu J, Du R, Ji G, Zhu L, Zhang J, Lv P, Cao J. Systematical characterization of GRF gene family in sorghum, and their potential functions in aphid resistance. Gene. 2022 Aug 20;836:146669. PMID: 35710084. DOI: 10.1016/j.gene.2022.146669. Read more

