Stable Protein, Starch and Amylose QTLs Identified for SC35 × RTx430 Sorghum Population
A persisting problem in the study of sorghum is altering grain quality without adversely affecting yield. Grain-quality traits like protein, starch and amylose content determine the end-use of sorghum crops. Approximately 70 percent of the dry weight of sorghum is due to starch content, while 12 percent is due to protein content. Amylose, one of the polysaccharides that compose starch, can comprise up to 30 percent of the starch in the plant, depending on the underlying genetics. In addition, mutations on the genes that regulate the storage proteins (kafirins). Understanding the genetics and conditions that produce sorghum crops geared towards specific end uses is of great interest to breeders and scientists.
In an effort to identity and map QTLs (quantitative trait loci) controlling protein, starch, and amylose content, and investigate the interaction with the environment in the production of sorghum cultivars for desired end-uses, Ayalew and colleagues from Kansas State University and the USDA-ARS studied a sorghum recombinant inbred line (RIL) population (RTx430 x SC35) grown in six different environments in Hays and Manhattan, Kansas. In total, seven protein, ten starch and ten amylose content QTLs were located. The researchers identified two QTL hotspots across multiple environments on chromosomes 1 and 2 which were associated with increased protein content. In addition to these, other protein content QTLs were located only in specific environments. For example, on chromosome 7, a QTL was detected in irrigated environments; this is consistent with the region where the Sobic.007G198600.1 gene, previously shown to be involved in stress response, growth and reproduction is located (Lechner et al., 2011). In contrast the QTLs identified for starch content were less consistent across environments. QTLs were identified on chromosomes 2, 3, 9 and 10, with the starch content QTL on the long arm of chromosome 2 co-locating with a protein content QTL, and presenting a future opportunity for selecting for simultaneous trait improvement. The scientists focused in on a specific 200-kb region of chromosome 10, which is known to be involved in amylose content, and identified 20 genes including the waxy (Wx) gene (Sobic.010G226000), which McIntyre et al.(2008) previously discovered helps regulate amylose production. The researchers further analyzed the candidate genes for all three grain qualities and their results indicated that the QTLs were enriched for transcription factors, including Cytochrome P450 and basic helix–loop–helix DNA binding protein, which regulate starch and protein accumulation in the grain.
The RILs in the study had large genetic variation for grain quality, which could be used to select for specific traits in line with the desired end-uses. The researchers also determined that a large proportion of the phenotypic variation was due to genetic variation (additive, dominant, and epistatic genes), which also supports the idea that grain quality could be improved in sorghum through genetic selection.
To complement breeding for yield gains in sorghum, identifying genomic regions that control grain quality would help develop high yielding nutritious sorghum for future changing climate. – Jagadish
SorghumBase Example:
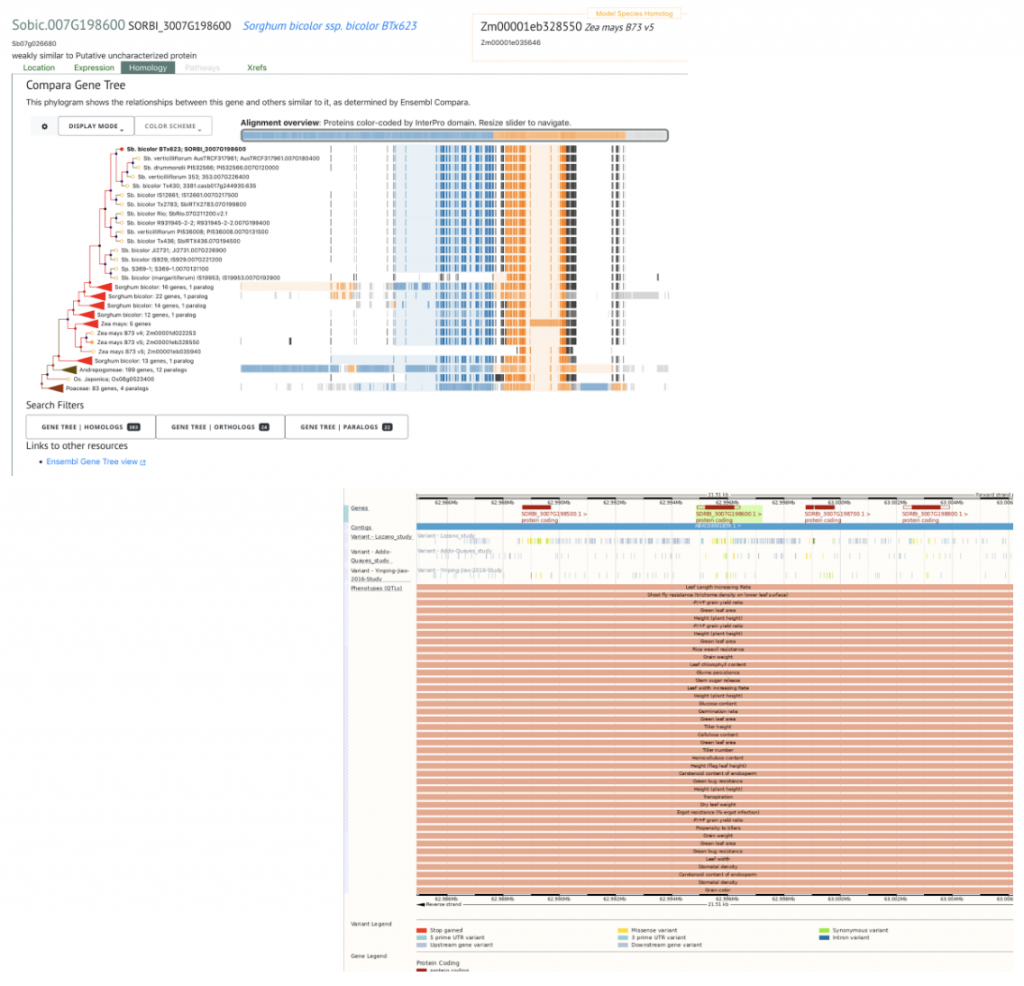
References:
Ayalew H, Peiris S, Chiluwal A, Kumar R, Tiwari M, Ostmeyer T, Bean S, Jagadish SVK. Stable sorghum grain quality QTL were identified using SC35 × RTx430 mapping population. Plant Genome. 2022 Jul 26:e20227. PMID: 35880472. DOI: 10.1002/tpg2.20227. Read more
Lechner E, Leonhardt N, Eisler H, Parmentier Y, Alioua M, Jacquet H, Leung J, Genschik P. MATH/BTB CRL3 receptors target the homeodomain-leucine zipper ATHB6 to modulate abscisic acid signaling. Dev Cell. 2011 Dec 13;21(6):1116-28. PMID: 22172674. DOI: 10.1016/j.devcel.2011.10.018. Read more
McIntyre CL, Drenth J, Gonzalez N, Henzell RG, Jordan DR. Molecular characterization of the waxy locus in sorghum. Genome. 2008 Jul;51(7):524-33. PMID: 18545276. DOI: 10.1139/G08-035. Read more


